Blog
Journal Club: Probing the Mechanisms of Triptolide-Induced Nephrotoxicity

At the second Journal Club session in our laboratory, held on April 10, 2026, we discussed the following paper:
Luo Y-M, Yang S-D, Wen M-Y, Wang B, Liu J-H, Li S-T, Li Y-Y, Cheng H, Zhao L-L, Li S-M, and Jiang J-J (2023)
Insights into the mechanisms of triptolide nephrotoxicity through network pharmacology-based analysis and RNA-seq.
Frontiers in Plant Science 14:1144583.
https://doi.org/10.3389/fpls.2023.1144583
Background
This paper focuses on triptolide (TPL), a natural compound isolated from Tripterygium wilfordii, a medicinal plant widely used in traditional Chinese medicine. Triptolide has been reported to possess a range of potent biological activities, including anti-inflammatory, immunosuppressive, and antitumor effects. At the same time, however, it is also known to cause adverse effects, including nephrotoxicity.
Because of this dual nature, as both a promising bioactive compound and a toxicological concern, triptolide has drawn considerable scientific interest. As a plant-derived compound with both pharmacological relevance and safety concerns, it made this paper a particularly compelling choice for our journal club.
The article was published in Frontiers in Plant Science, a journal that covers a broad range of topics such as plant genetics, physiology, ecology, and biotechnology. Since triptolide is derived from a medicinal plant and the study explores its toxic mechanism at the molecular level, the work also fits well within the journal’s broader scope.
Main Findings of the Study
To investigate the molecular basis of TPL-induced nephrotoxicity, the authors combined network pharmacology analysis with RNA-seq. Using this integrative approach, they proposed that triptolide-induced kidney injury may involve both c-Jun-centered inflammatory signaling and circadian rhythm disruption mediated by Per1.
A major strength of the study is that it does not attempt to explain toxicity through a single molecule alone. Instead, it uses a systems-level approach to generate mechanistic hypotheses by integrating multiple pathways and datasets. This is particularly valuable when studying compounds whose toxic effects are recognized, but whose overall molecular mechanism remains incompletely understood.
Figure
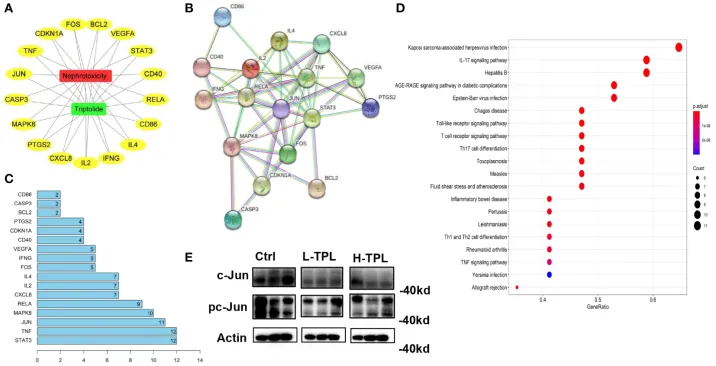
Figure. Identification of target genes associated with triptolide-induced nephrotoxicity by network pharmacology analysis (reproduced from Figure 2 of the original article).
Luo Y-M, Yang S-D, Wen M-Y, et al. Insights into the mechanisms of triptolide nephrotoxicity through network pharmacology-based analysis and RNA-seq. Frontiers in Plant Science. 2023;14:1144583. DOI: 10.3389/fpls.2023.1144583. Licensed under CC BY.
A Question Raised During the Discussion
One of the questions raised during the session concerned the interpretation of the circadian rhythm-related findings:
If Per1-mediated circadian disruption is involved, could the results have been influenced by sampling time, given that circadian genes naturally fluctuate over the course of the day?
This led to an important discussion. Because circadian rhythm-related genes are inherently time-dependent, it is difficult to completely rule out the possibility that diurnal variation contributed to some of the observed expression changes. At the same time, this discussion reinforced an important experimental point: when studying circadian pathways, it is essential to standardize sample collection time across experimental groups as strictly as possible.
Why I Chose This Paper
I am currently studying the toxic mechanism of a novel compound, so I was particularly interested in this paper’s strategy for investigating a substance whose mechanism has not yet been fully clarified. The combination of RNA-seq and network-based analysis to identify upstream regulators and related pathways was especially appealing.
What stood out to me was the way the authors used publicly available large-scale datasets to generate mechanistic hypotheses for a biologically active compound whose overall toxic mechanism remains unclear. I felt that this framework could also be applicable to my own research and could provide valuable clues for organizing the molecular networks underlying toxic responses.
Take-Home Message
This paper provided an instructive example of how integrative omics approaches can help uncover the mechanisms of toxic injury. Rather than treating toxicity as the consequence of a single pathway, the study approached it as a complex biological response involving inflammation, circadian regulation, and multiple interacting signals.
For our lab, the paper served as a useful reminder that combining RNA-seq with network analysis can be a powerful strategy for generating mechanistic insight into compounds with unresolved toxicological profiles.
Reference
Luo Y-M, Yang S-D, Wen M-Y, Wang B, Liu J-H, Li S-T, Li Y-Y, Cheng H, Zhao L-L, Li S-M, and Jiang J-J (2023).
Insights into the mechanisms of triptolide nephrotoxicity through network pharmacology-based analysis and RNA-seq.
Frontiers in Plant Science. 14:1144583.
https://doi.org/10.3389/fpls.2023.1144583
Latest Blog
Category
Archive
- 2026-4 (3)
- 2025-7 (6)
- 2025-6 (2)
- 2025-5 (2)
- 2025-4 (4)
- 2025-3 (4)
- 2025-2 (1)
- 2025-1 (2)
- 2024-12 (1)
- 2024-11 (3)
- 2024-10 (5)
- 2024-9 (1)
- 2024-8 (1)
- 2024-7 (4)
- 2024-6 (1)
- 2024-4 (1)
- 2024-2 (1)
- 2024-1 (1)
- 2023-11 (1)
- 2023-10 (1)
- 2023-6 (1)
- 2023-4 (1)
- 2023-3 (1)
- 2022-12 (1)
- 2022-11 (1)
- 2022-9 (1)
- 2022-6 (1)
- 2022-3 (3)
- 2022-1 (1)
- 2021-12 (3)
- 2021-11 (1)
- 2021-10 (1)
- 2021-9 (1)
- 2021-8 (1)
- 2021-7 (3)